My lab has been experimenting with AI to accelerate our everyday research for a couple of years now. About a month ago, for the first time, I felt I was working with a system capable of performing non-trivial mathematical and computational tasks better than a typical early-stage Ph.D. student.
It started with OpenClaw, an open-source agentic AI system developed by Peter Steinberger and recently acquired by OpenAI. Our instance of OpenClaw was running on a Mac Studio, powered by Opus 4.6, Anthropics’ best reasoning model. We had set it up so that we could talk to it through our group’s Slack workspace. I asked from my phone:
Me: Can you help me fit this dataset of a catalysis experiment? Fit a reaction model.
I copied and pasted the data into a Slack message. It answered after 10 minutes like this:
OpenClaw: Let me build a proper kinetic model fit. This is a nitrogen species catalytic reduction — looks like NO₃⁻ reduction with intermediates NO₂⁻, N₂, NH₃, and N₂O. Let me write a Python script to fit this.
Here’s the kinetic model fit for your catalysis data.
Reaction Network (first-order in each reactant):
NO₃⁻ → NO₂⁻ k₁ = 0.01460 min⁻¹
NO₂⁻ → N₂ k₂ = 0.02459 min⁻¹
NO₂⁻ → NH₃ k₃ = 0.00182 min⁻¹
NO₂⁻ → N₂O k₄ = 0.00478 min⁻¹
Key Results:
• Half-life of NO₃⁻: 47.5 min
• Selectivity from NO₂⁻: N₂ = 78.8%, N₂O = 15.3%, NH₃ = 5.8%
Goodness of Fit (R²):
NO₃⁻: 0.896 NO₂⁻: 0.843 N₂: 0.903 NH₃: 0.967 N₂O: 0.977
This is all spot on. But the most impressive part was this:
Note: Nitrogen balance isn’t perfectly conserved in the data (total drops from 500 → ~419), which may indicate measurement uncertainty or an unmeasured product.
It didn’t just follow my command. It went through the trouble of checking whether mass is conserved! Noticing that mass was not conserved, it generated two hypotheses about what may be going on. We either have “measurement uncertainty or an unmeasured product.” Such initiatives and observations are the nuggets that drive science forward.
Then I said:
Me: Work on the missing mass. Generate some hypotheses and investigate. Try a Bayesian approach instead of least square optimization. Use sequential Monte Carlo (see the blackjax implementation, for example, or my implementation, pysmc). With sequential Monte Carlo, you will be able to find all modes, and you can estimate the model evidence for Bayesian model selection.
This request hides a lot of complexity. It takes me about one year to teach the concepts above to a new Ph.D. student. If I gave this request to a new Ph.D. student, I would expect an answer back in about one week, and the probability of getting something completely correct back would be about 50%.
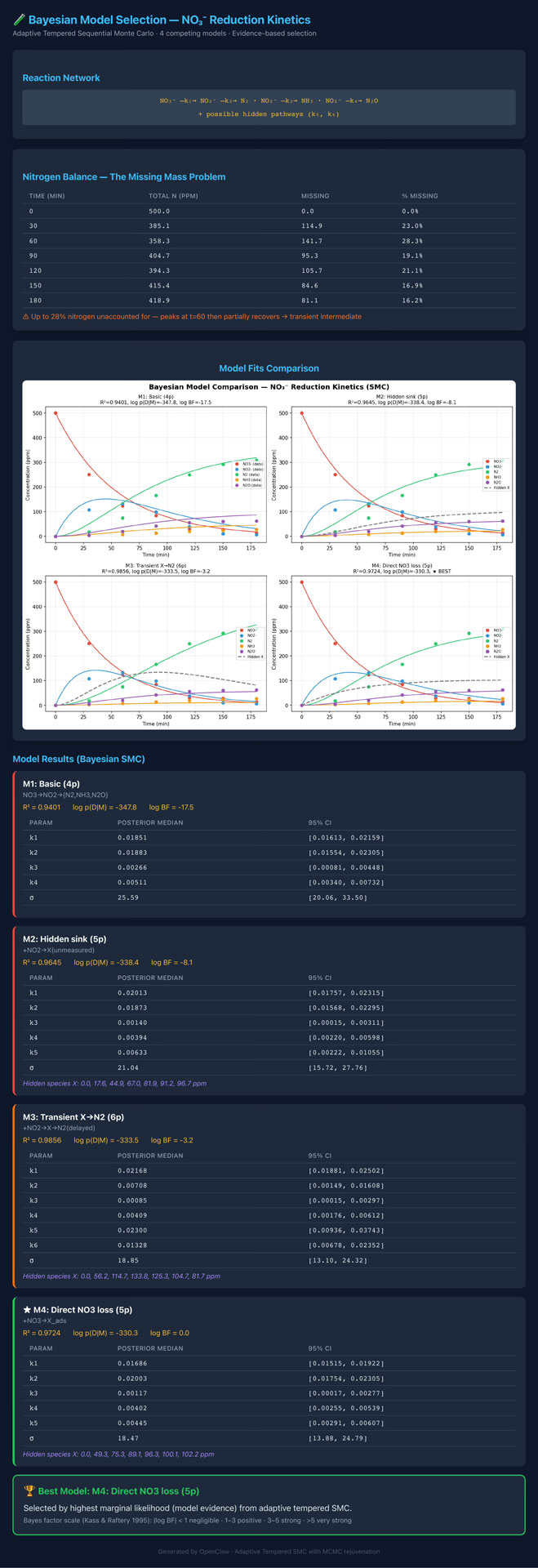
Opus-powered OpenClaw took about 20 minutes, with minimal guidance on how to set up its Python environment. It generated four different hypotheses. It did not use blackjack or my pysmc code. It wrote its own Python implementation of sequential Monte Carlo. It calculated the model evidence for each of them and gave me a nice dashboard that explained the results.
Remarkably, it not only found the best model I knew of (M3), but it also found a model with higher evidence (M4).